Directed Evolution and the AutoDiMe Method
Comparison of Directed Evolution and AutoDiMe
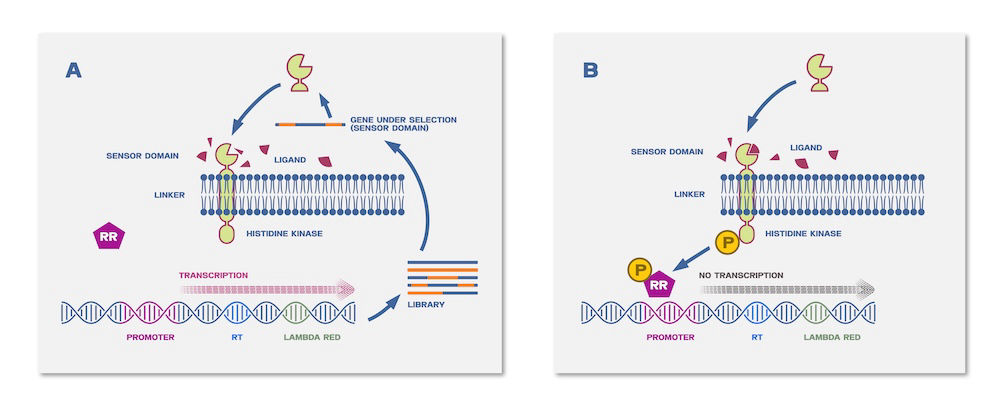
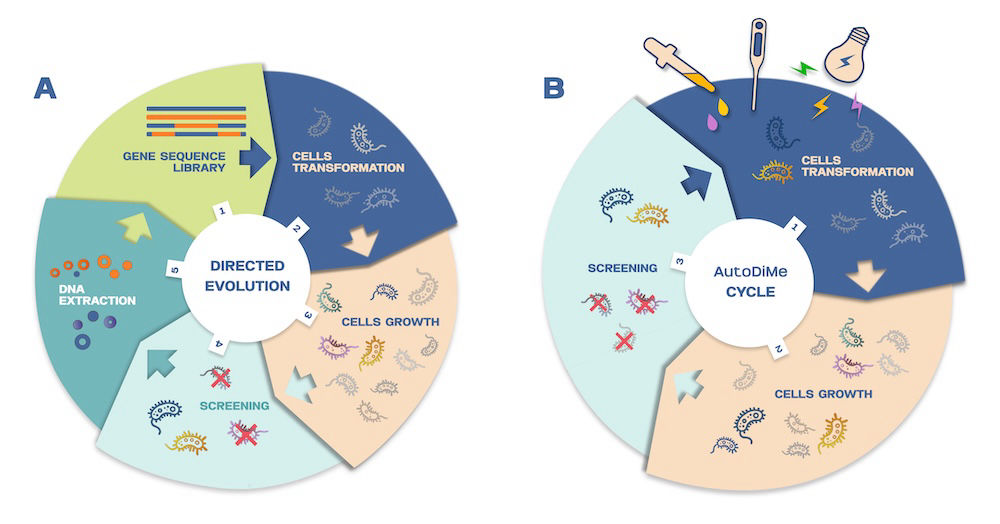
This figure compares classical directed evolution with the AutoDiMe method. In classical approaches, selected variants are isolated, and their DNA is extracted, recombined in vitro, and reintroduced into cells in iterative cycles. In contrast, AutoDiMe enables diversification directly in vivo through recombination of homologous, reverse-transcribed sequences. After initial setup, the process can proceed autonomously, allowing continuous generation and selection of variants within the cell population. External control (e.g., temperature shifts) can regulate or stop the process when desired outcomes are achieved.